Structure of the T. vaginalis MULEs. Putative terminal inverted repeats... | Download Scientific Diagram

U-type exchange is the most frequent mechanism for inverted duplication with terminal deletion rearrangements | Journal of Medical Genetics

Terminal inverted repeats sequence of DPLT1-4 transposons and related... | Download Scientific Diagram

Minos-derived vectors. Minos inverted terminal repeats are shown as... | Download Scientific Diagram
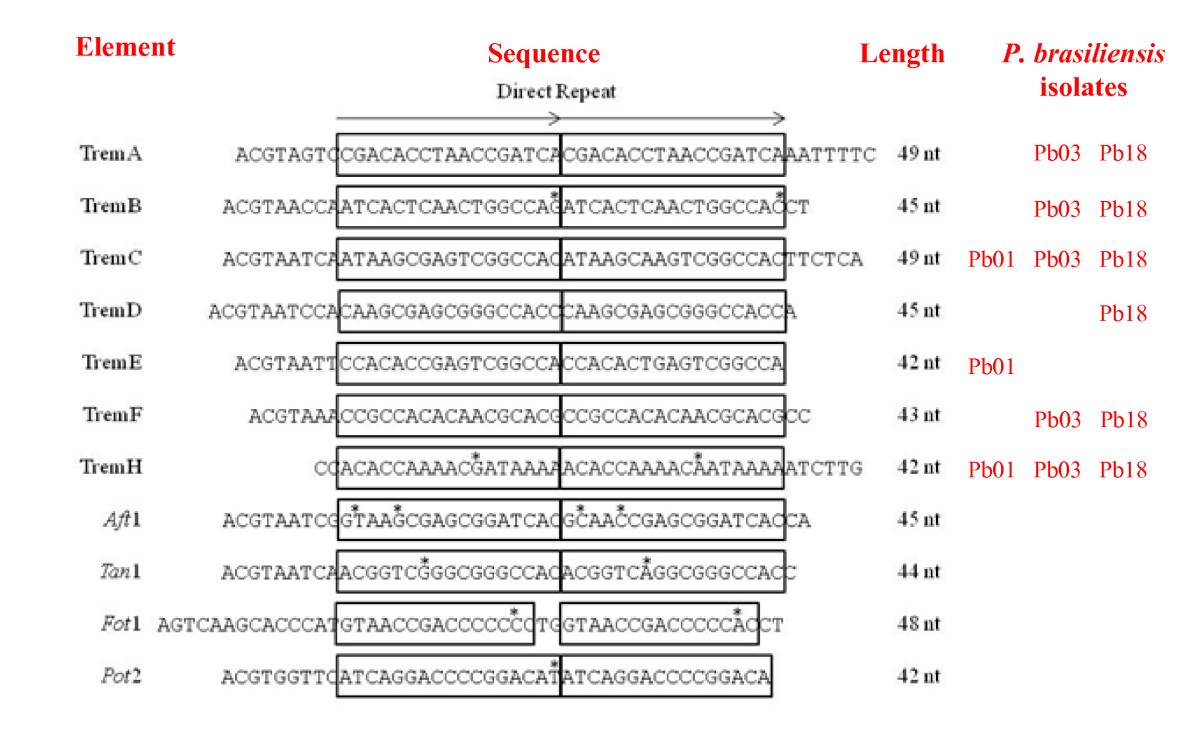
Identification and characterization of Tc1/mariner-like DNA transposons in genomes of the pathogenic fungi of the Paracoccidioides species complex | BMC Genomics | Full Text
PLOS ONE: Insertion Sequence Inversions Mediated by Ectopic Recombination between Terminal Inverted Repeats

Models for terminal inverted repeat (TIR) expansion and reduction. (a)... | Download Scientific Diagram
PLOS ONE: Mobility and Generation of Mosaic Non-Autonomous Transposons by Tn3-Derived Inverted-Repeat Miniature Elements (TIMEs)
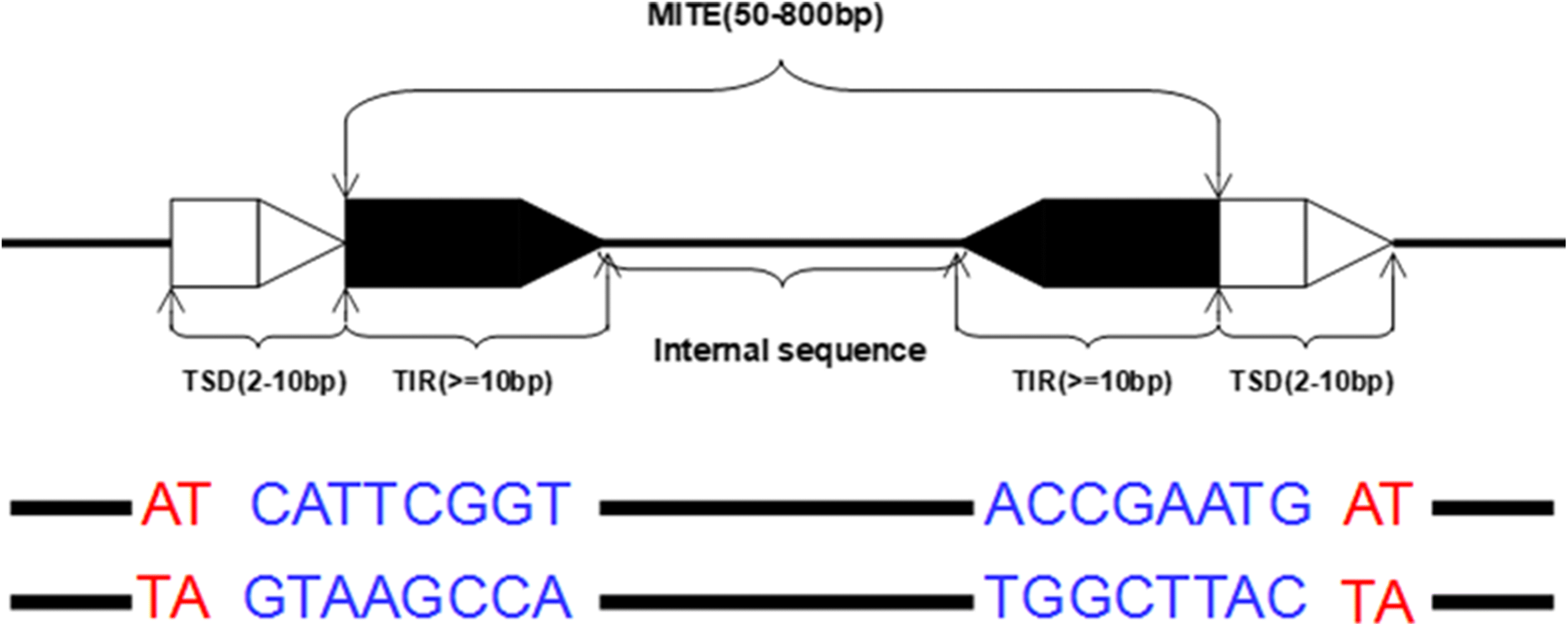
MiteFinderII: a novel tool to identify miniature inverted-repeat transposable elements hidden in eukaryotic genomes | BMC Medical Genomics | Full Text

detectMITE: A novel approach to detect miniature inverted repeat transposable elements in genomes. - Abstract - Europe PMC
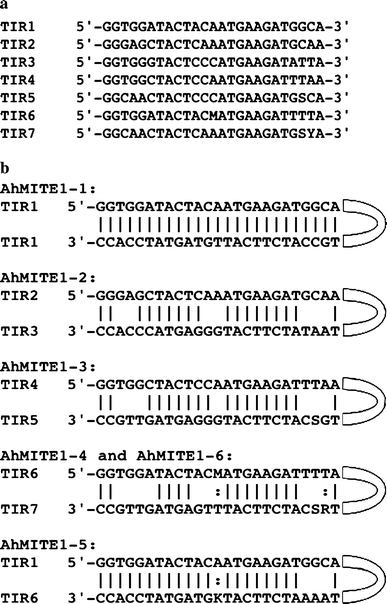
Characterization of active miniature inverted-repeat transposable elements in the peanut genome | SpringerLink
![PDF] Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar PDF] Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/c29fd938f405ed8516c10b9bdaa27f988b29dd31/2-Figure1-1.png)
PDF] Structure-function analysis of the inverted terminal repeats of the sleeping beauty transposon. | Semantic Scholar
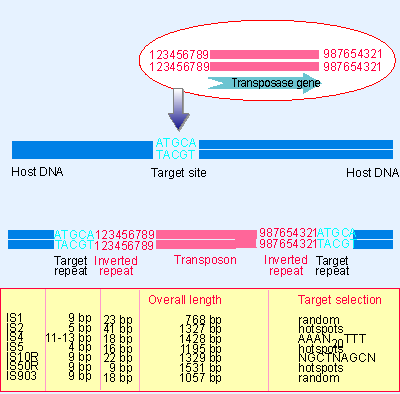
GENES IV INDEX > Transposons Transposons 15.2 Insertion sequences are simple transposition modules Key terms defined in this section Direct repeats are identical (or related) sequences present in two or more copies in the same orientation in the same ...